Advanced Topics
This guide covers advanced statistical concepts, experimental designs, and optimization techniques for sophisticated users of GeoStep.
System Architecture Overview
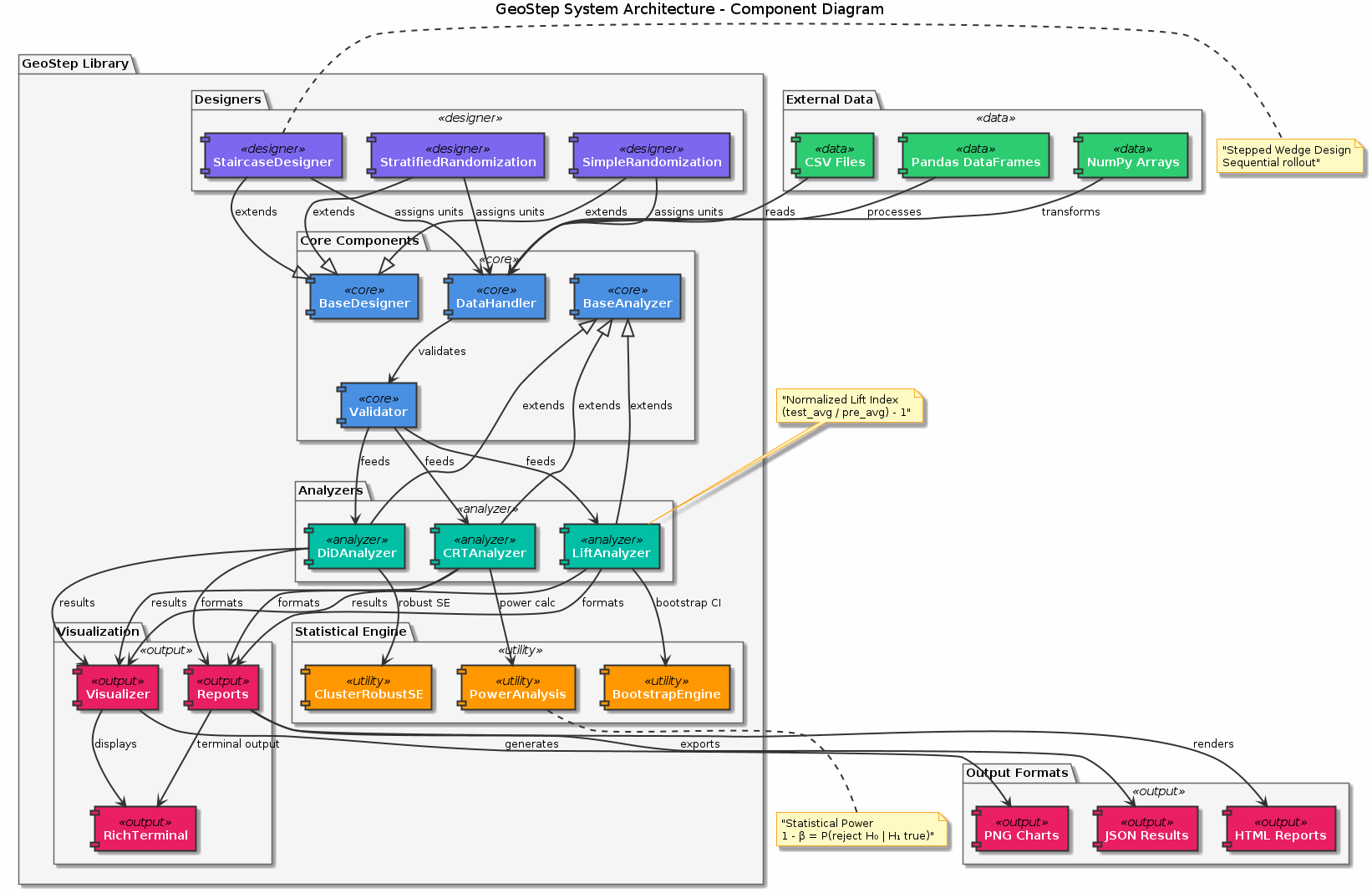 Figure: Component architecture of the GeoStep library showing the modular design with analyzers (LiftAnalyzer, DiDAnalyzer, CRTAnalyzer), designers (SimpleRandomization, Stratified, Staircase), and supporting utilities for data handling, visualization, and reporting.
Figure: Component architecture of the GeoStep library showing the modular design with analyzers (LiftAnalyzer, DiDAnalyzer, CRTAnalyzer), designers (SimpleRandomization, Stratified, Staircase), and supporting utilities for data handling, visualization, and reporting.
🧮 Statistical Methodology Deep Dive
Randomized Controlled Trials (RCTs) Theory
Fundamental Assumptions
SUTVA (Stable Unit Treatment Value Assumption): Treatment of one unit doesn’t affect outcomes of other units
Ignorability: Treatment assignment is independent of potential outcomes
Positivity: All units have non-zero probability of receiving any treatment
Mathematical Framework
For geographic unit $i$ in time period $t$:
Y_it(1) = potential outcome under treatment
Y_it(0) = potential outcome under control
τ_i = Y_it(1) - Y_it(0) = individual treatment effect
ATE = E[τ_i] = average treatment effect
The randomization ensures:
E[Y_it(1)|T_i = 1] = E[Y_it(1)|T_i = 0]
E[Y_it(0)|T_i = 1] = E[Y_it(0)|T_i = 0]
Normalization Method Details
GeoStep uses a normalized lift index approach:
Pre-period normalization:
baseline_i = mean(Y_it for t in pre_period)
Expected performance:
expected_it = baseline_i * seasonal_factor_t
Lift index calculation:
lift_index_i = (actual_i - expected_i) / expected_i
Treatment effect estimation:
ATE = mean(lift_index_treatment) - mean(lift_index_control)
This approach is robust to:
Scale differences between geographic units
Seasonal patterns in the data
Natural growth trends
Power Analysis Mathematics
Statistical power is the probability of detecting a true effect:
Power = P(reject H0 | H1 is true)
= P(|t-statistic| > critical_value | true_effect ≠ 0)
For a two-sample t-test with effect size δ:
Power = Φ(δ√(n/2) - z_α/2) + Φ(-δ√(n/2) - z_α/2)
Where:
Φ= standard normal CDFδ= standardized effect sizen= sample size per groupz_α/2= critical value for significance level α
🏗️ Advanced Experimental Designs
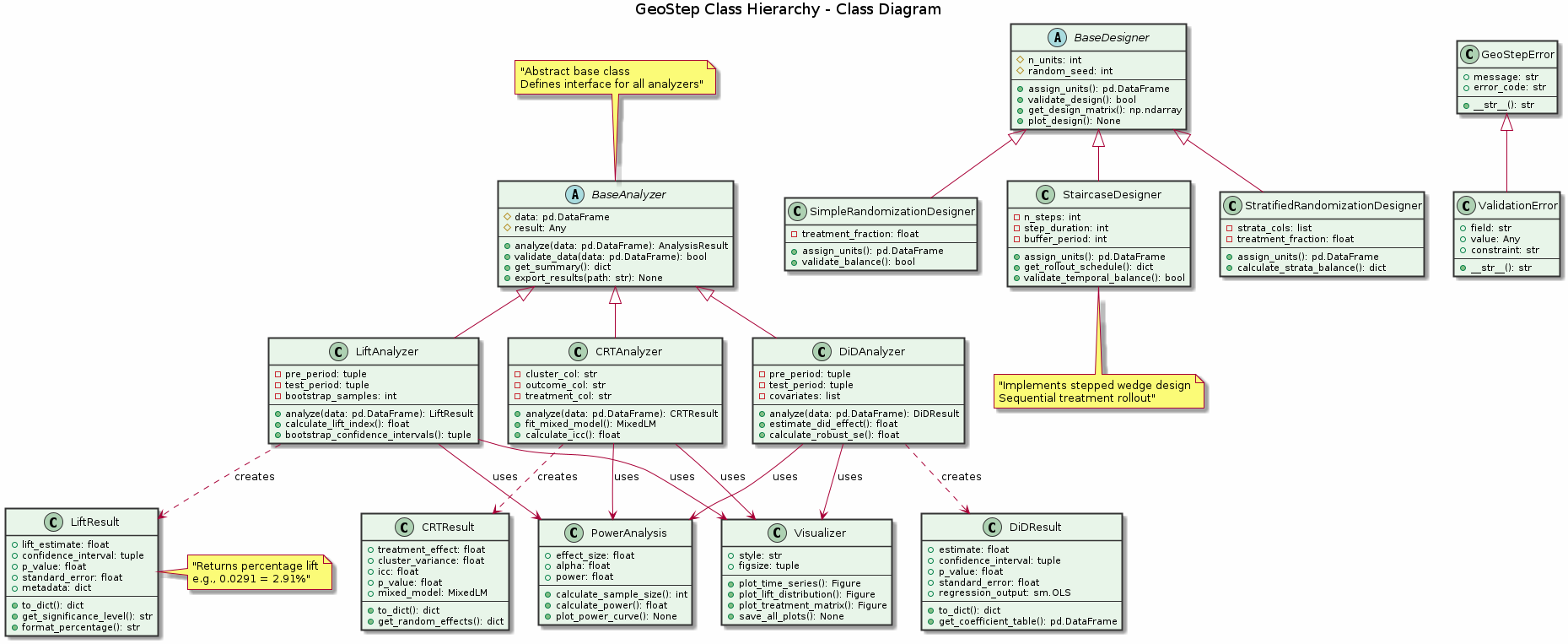 Figure: Object-oriented class hierarchy showing inheritance relationships. BaseAnalyzer and BaseDesigner serve as abstract base classes, with concrete implementations for different analysis methods and experimental designs.
Figure: Object-oriented class hierarchy showing inheritance relationships. BaseAnalyzer and BaseDesigner serve as abstract base classes, with concrete implementations for different analysis methods and experimental designs.
Stepped-Wedge Trials
Design Matrix
For a stepped-wedge with K clusters and T time periods:
import numpy as np
def create_stepped_wedge_matrix(K, T, step_size=1):
"""
Creates a stepped-wedge design matrix.
Parameters:
K: Number of clusters
T: Number of time periods
step_size: Number of clusters switching per step
"""
design = np.zeros((K, T))
# Calculate switch times
switch_times = np.arange(1, T, T//K)
for i, switch_time in enumerate(switch_times):
cluster_start = i * step_size
cluster_end = min((i + 1) * step_size, K)
design[cluster_start:cluster_end, switch_time:] = 1
return design
Analysis Model
Mixed-effects model for stepped-wedge:
Y_ijt = β0 + β1*X_jt + α_j + γ_t + ε_ijt
Where:
Y_ijt= outcome for individual i in cluster j at time tX_jt= treatment indicator (0/1)α_j= cluster random effectγ_t= time fixed effectβ1= treatment effect (parameter of interest)
Staircase Design Optimization
Optimal Allocation
For a staircase design, the optimal number of pre/post periods balances:
Statistical power (more periods = better)
Operational cost (fewer periods = cheaper)
def optimize_staircase_design(total_budget, cost_per_period,
effect_size, alpha=0.05, power=0.8):
"""
Find optimal pre/post period allocation for staircase design.
"""
best_power = 0
best_design = None
for pre_periods in range(2, 10):
for post_periods in range(2, 10):
total_cost = (pre_periods + post_periods) * cost_per_period
if total_cost <= total_budget:
# Calculate power for this design
power = calculate_staircase_power(
pre_periods, post_periods, effect_size, alpha
)
if power > best_power:
best_power = power
best_design = (pre_periods, post_periods)
return best_design, best_power
Cluster Randomized Trials (CRTs)
Intracluster Correlation (ICC)
The ICC measures similarity within clusters:
ICC = σ²_between / (σ²_between + σ²_within)
Higher ICC reduces effective sample size:
Design_Effect = 1 + (m - 1) * ICC
Effective_n = n / Design_Effect
Where m = average cluster size.
Sample Size Calculation
For CRTs, required number of clusters:
def calculate_crt_sample_size(effect_size, icc, cluster_size,
alpha=0.05, power=0.8):
"""
Calculate required number of clusters for CRT.
"""
from scipy import stats
z_alpha = stats.norm.ppf(1 - alpha/2)
z_beta = stats.norm.ppf(power)
design_effect = 1 + (cluster_size - 1) * icc
n_clusters = (2 * design_effect * (z_alpha + z_beta)**2) / \
(cluster_size * effect_size**2)
return int(np.ceil(n_clusters))
⚡ Performance Optimization
Large-Scale Experiments
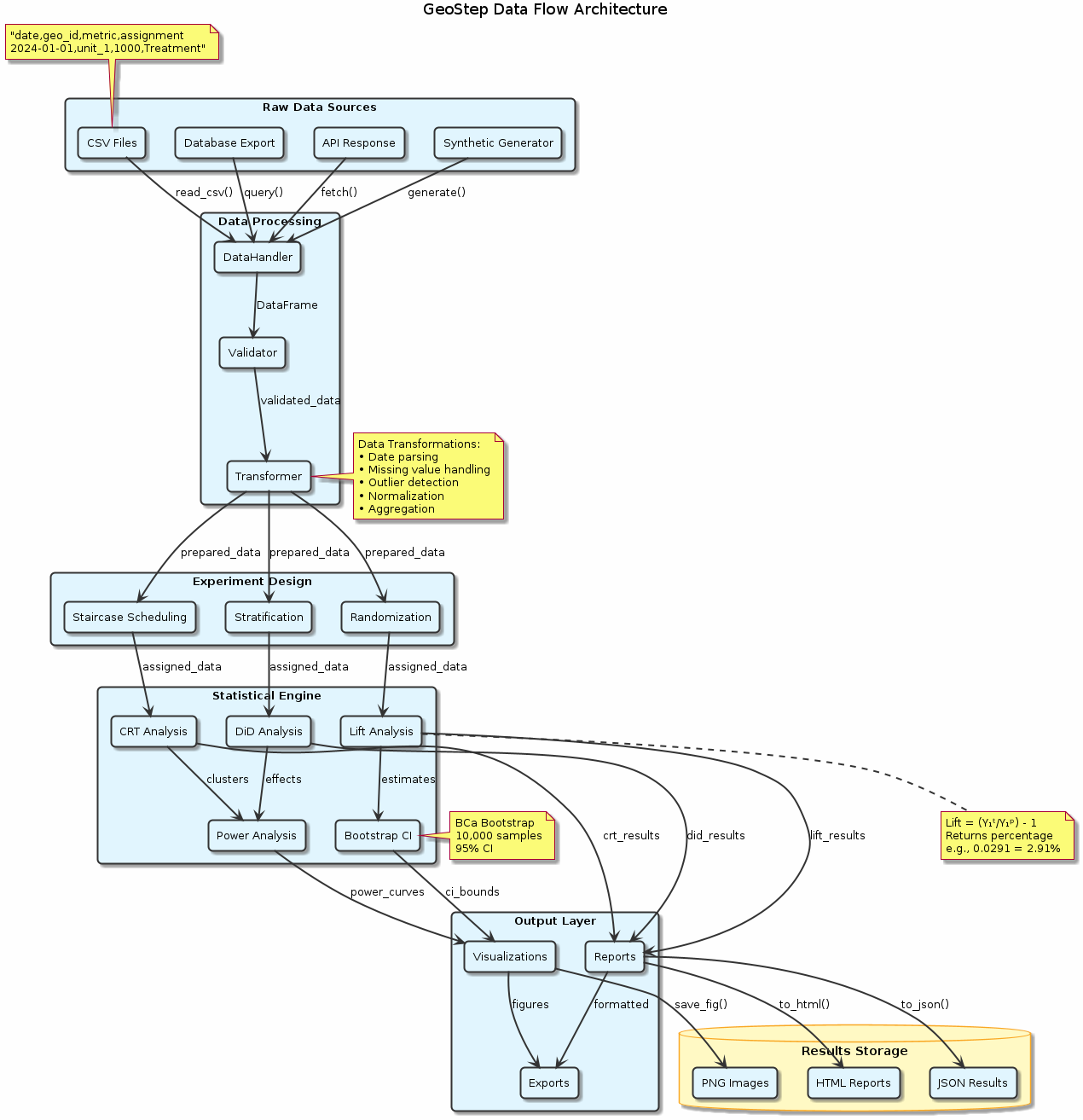 Figure: End-to-end data flow from raw sources through processing pipeline to final outputs. The architecture supports multiple data formats and provides both programmatic and visual outputs for comprehensive analysis.
Figure: End-to-end data flow from raw sources through processing pipeline to final outputs. The architecture supports multiple data formats and provides both programmatic and visual outputs for comprehensive analysis.
Data Sampling for Power Analysis
For datasets with millions of rows:
def stratified_sample_for_power(df, geo_col, strata_col,
sample_size=50000):
"""
Create stratified sample for power analysis.
"""
# Calculate strata proportions
strata_props = df[strata_col].value_counts(normalize=True)
sampled_data = []
for stratum, prop in strata_props.items():
stratum_data = df[df[strata_col] == stratum]
stratum_sample_size = int(sample_size * prop)
if len(stratum_data) > stratum_sample_size:
stratum_sample = stratum_data.sample(
n=stratum_sample_size,
random_state=42
)
else:
stratum_sample = stratum_data
sampled_data.append(stratum_sample)
return pd.concat(sampled_data, ignore_index=True)
Parallel Processing Configuration
import os
from joblib import Parallel, delayed
# Configure parallel processing
os.environ['NUMBA_NUM_THREADS'] = '8' # Limit CPU usage
os.environ['OMP_NUM_THREADS'] = '8'
# Custom parallel power analysis
def parallel_power_analysis(historical_data, effect_sizes,
test_weeks_list, n_jobs=4):
"""
Run power analysis with custom parallelization.
"""
def single_scenario(effect, weeks):
return run_single_power_scenario(
historical_data, effect, weeks
)
scenarios = [(e, w) for e in effect_sizes
for w in test_weeks_list]
results = Parallel(n_jobs=n_jobs)(
delayed(single_scenario)(effect, weeks)
for effect, weeks in scenarios
)
return pd.DataFrame(results)
Memory Optimization
Efficient Data Types
def optimize_datatypes(df):
"""
Optimize DataFrame memory usage.
"""
# Convert object columns to category
for col in df.select_dtypes(include=['object']):
if df[col].nunique() < 0.5 * len(df):
df[col] = df[col].astype('category')
# Downcast numeric types
for col in df.select_dtypes(include=['int64']):
df[col] = pd.to_numeric(df[col], downcast='integer')
for col in df.select_dtypes(include=['float64']):
df[col] = pd.to_numeric(df[col], downcast='float')
return df
🔬 Advanced Analysis Techniques
Sensitivity Analysis
Robustness Checks
def sensitivity_analysis(df, geo_col, assignment_col,
date_col, kpi_col, test_start):
"""
Perform sensitivity analysis.
"""
results = {}
# 1. Different pre-period lengths
for pre_weeks in [4, 6, 8, 10, 12]:
pre_end = pd.to_datetime(test_start) - pd.Timedelta(days=1)
pre_start = pre_end - pd.Timedelta(weeks=pre_weeks)
subset_data = df[df[date_col] >= pre_start]
result = run_primary_analysis(subset_data, ...)
results[f'pre_weeks_{pre_weeks}'] = result
# 2. Outlier removal
for percentile in [95, 97.5, 99]:
threshold = df[kpi_col].quantile(percentile/100)
clean_data = df[df[kpi_col] <= threshold]
result = run_primary_analysis(clean_data, ...)
results[f'outlier_removal_{percentile}'] = result
# 3. Different analysis methods
results['primary'] = run_primary_analysis(df, ...)
results['did'] = run_did_analysis(df, ...)
return results
Bayesian Analysis
Bayesian Treatment Effect Estimation
import pymc as pm
import arviz as az
def bayesian_rct_analysis(treatment_data, control_data):
"""
Bayesian analysis of RCT results.
"""
with pm.Model() as model:
# Priors
mu_control = pm.Normal('mu_control', mu=0, sigma=1)
mu_treatment = pm.Normal('mu_treatment', mu=0, sigma=1)
sigma = pm.HalfNormal('sigma', sigma=1)
# Likelihood
control_obs = pm.Normal('control_obs',
mu=mu_control,
sigma=sigma,
observed=control_data)
treatment_obs = pm.Normal('treatment_obs',
mu=mu_treatment,
sigma=sigma,
observed=treatment_data)
# Treatment effect
treatment_effect = pm.Deterministic(
'treatment_effect',
mu_treatment - mu_control
)
# Sampling
trace = pm.sample(2000, tune=1000,
return_inferencedata=True)
return trace
Causal Inference Extensions
Instrumental Variables
For cases with non-compliance:
def instrumental_variables_analysis(df, instrument_col,
treatment_col, outcome_col):
"""
Two-stage least squares for IV analysis.
"""
from sklearn.linear_model import LinearRegression
# First stage: instrument -> treatment
X_first = df[[instrument_col]]
y_first = df[treatment_col]
first_stage = LinearRegression().fit(X_first, y_first)
# Predicted treatment
treatment_hat = first_stage.predict(X_first)
# Second stage: predicted treatment -> outcome
X_second = treatment_hat.reshape(-1, 1)
y_second = df[outcome_col]
second_stage = LinearRegression().fit(X_second, y_second)
# IV estimate
iv_estimate = second_stage.coef_[0]
return {
'iv_estimate': iv_estimate,
'first_stage_r2': first_stage.score(X_first, y_first),
'weak_instrument_test': first_stage_r2 > 0.1
}
🚀 Production Deployment
Automated Experiment Pipeline
Configuration Management
from dataclasses import dataclass
from typing import List, Optional
import yaml
@dataclass
class ExperimentConfig:
"""Configuration for automated experiments."""
name: str
geo_col: str
date_col: str
kpi_col: str
treatment_description: str
expected_effect_size: float
test_duration_weeks: int
randomization_method: str = 'simple'
stratification_vars: Optional[List[str]] = None
power_threshold: float = 0.8
@classmethod
def from_yaml(cls, yaml_path: str):
with open(yaml_path, 'r') as f:
config_dict = yaml.safe_load(f)
return cls(**config_dict)
Experiment Orchestration
class ExperimentOrchestrator:
"""Orchestrates end-to-end experiment workflow."""
def __init__(self, config: ExperimentConfig):
self.config = config
self.results = {}
def run_full_pipeline(self, data_path: str):
"""Run complete experiment pipeline."""
# 1. Load and validate data
data = self.load_and_validate_data(data_path)
# 2. Power analysis
power_results = self.run_power_analysis(data)
# 3. Check if experiment is viable
if not self.is_experiment_viable(power_results):
raise ValueError("Experiment not viable - insufficient power")
# 4. Randomization
assigned_data = self.randomize_units(data)
# 5. Balance check
balance_results = self.check_balance(assigned_data)
# 6. Generate experiment plan
self.generate_experiment_plan(assigned_data)
return {
'power_analysis': power_results,
'balance_check': balance_results,
'assignment': assigned_data,
'experiment_ready': True
}
Monitoring and Alerting
Real-time Monitoring
def monitor_experiment_execution(experiment_id: str,
data_source: str):
"""
Monitor experiment execution in real-time.
"""
import time
from datetime import datetime, timedelta
while True:
# Check data freshness
latest_data = get_latest_data(data_source)
data_lag = datetime.now() - latest_data['date'].max()
if data_lag > timedelta(days=2):
send_alert(f"Data lag detected: {data_lag}")
# Check treatment delivery
treatment_delivery = check_treatment_delivery(
experiment_id, latest_data
)
if treatment_delivery['coverage'] < 0.95:
send_alert(f"Low treatment coverage: {treatment_delivery}")
# Check for contamination
contamination = check_contamination(latest_data)
if contamination['spillover_detected']:
send_alert(f"Contamination detected: {contamination}")
time.sleep(3600) # Check hourly
📊 Advanced Reporting
Interactive Dashboards
import plotly.graph_objects as go
from plotly.subplots import make_subplots
def create_interactive_dashboard(experiment_results):
"""
Create interactive dashboard for experiment results.
"""
fig = make_subplots(
rows=2, cols=2,
subplot_titles=('Power Analysis', 'Time Series',
'Lift Distribution', 'Balance Check'),
specs=[[{"type": "heatmap"}, {"type": "scatter"}],
[{"type": "histogram"}, {"type": "bar"}]]
)
# Power analysis heatmap
fig.add_trace(
go.Heatmap(
z=experiment_results['power_matrix'],
colorscale='Viridis'
),
row=1, col=1
)
# Time series
fig.add_trace(
go.Scatter(
x=experiment_results['dates'],
y=experiment_results['treatment_trend'],
name='Treatment'
),
row=1, col=2
)
# Add more plots...
fig.update_layout(
title="Experiment Results Dashboard",
height=800
)
return fig
This advanced topics guide provides the mathematical foundation and sophisticated techniques needed for complex experimental designs and large-scale implementations of GeoStep.